Zinc »
PDB 2n9o-2nwo »
2n9p »
Zinc in PDB 2n9p: Solution Structure of RNF126 N-Terminal Zinc Finger Domain in Complex with BAG6 Ubiquitin-Like Domain
Zinc Binding Sites:
The binding sites of Zinc atom in the Solution Structure of RNF126 N-Terminal Zinc Finger Domain in Complex with BAG6 Ubiquitin-Like Domain
(pdb code 2n9p). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total only one binding site of Zinc was determined in the Solution Structure of RNF126 N-Terminal Zinc Finger Domain in Complex with BAG6 Ubiquitin-Like Domain, PDB code: 2n9p:
In total only one binding site of Zinc was determined in the Solution Structure of RNF126 N-Terminal Zinc Finger Domain in Complex with BAG6 Ubiquitin-Like Domain, PDB code: 2n9p:
Zinc binding site 1 out of 1 in 2n9p
Go back to
Zinc binding site 1 out
of 1 in the Solution Structure of RNF126 N-Terminal Zinc Finger Domain in Complex with BAG6 Ubiquitin-Like Domain
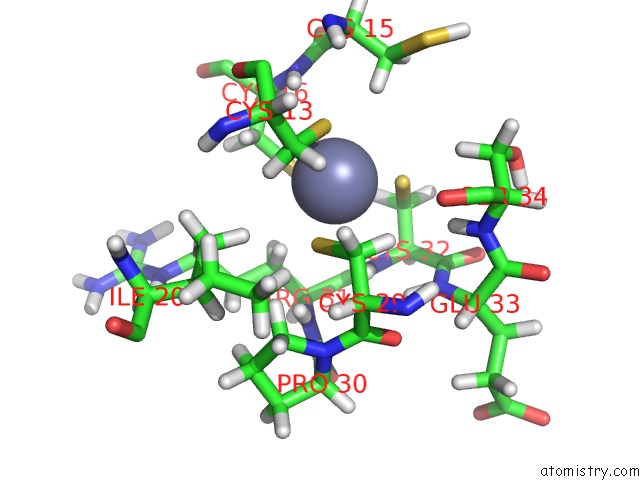
Mono view
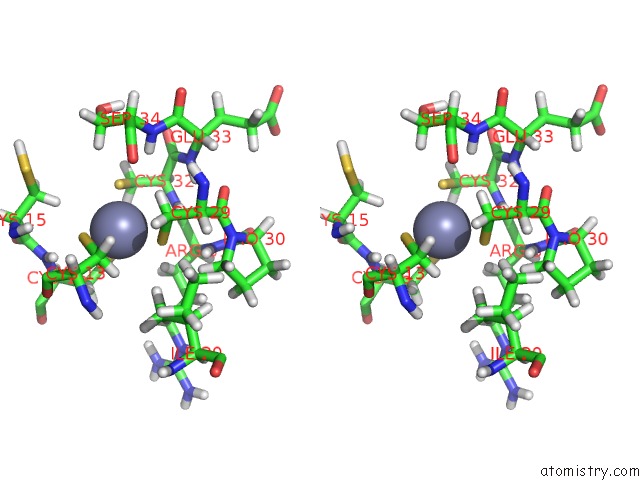
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Solution Structure of RNF126 N-Terminal Zinc Finger Domain in Complex with BAG6 Ubiquitin-Like Domain within 5.0Å range:
|
Reference:
E.M.Krysztofinska,
S.Martinez-Lumbreras,
A.Thapaliya,
N.J.Evans,
S.High,
R.L.Isaacson.
Structural and Functional Insights Into the E3 Ligase, RNF126. Sci Rep V. 6 26433 2016.
ISSN: ESSN 2045-2322
PubMed: 27193484
DOI: 10.1038/SREP26433
Page generated: Thu Oct 17 02:13:04 2024
ISSN: ESSN 2045-2322
PubMed: 27193484
DOI: 10.1038/SREP26433
Last articles
Zn in 9JYWZn in 9IR4
Zn in 9IR3
Zn in 9GMX
Zn in 9GMW
Zn in 9JEJ
Zn in 9ERF
Zn in 9ERE
Zn in 9EGV
Zn in 9EGW