Zinc in PDB 7nad: State E2 Nucleolar 60S Ribosomal Biogenesis Intermediate - SPB4 Local Refinement Model
Zinc Binding Sites:
The binding sites of Zinc atom in the State E2 Nucleolar 60S Ribosomal Biogenesis Intermediate - SPB4 Local Refinement Model
(pdb code 7nad). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total only one binding site of Zinc was determined in the State E2 Nucleolar 60S Ribosomal Biogenesis Intermediate - SPB4 Local Refinement Model, PDB code: 7nad:
In total only one binding site of Zinc was determined in the State E2 Nucleolar 60S Ribosomal Biogenesis Intermediate - SPB4 Local Refinement Model, PDB code: 7nad:
Zinc binding site 1 out of 1 in 7nad
Go back to
Zinc binding site 1 out
of 1 in the State E2 Nucleolar 60S Ribosomal Biogenesis Intermediate - SPB4 Local Refinement Model
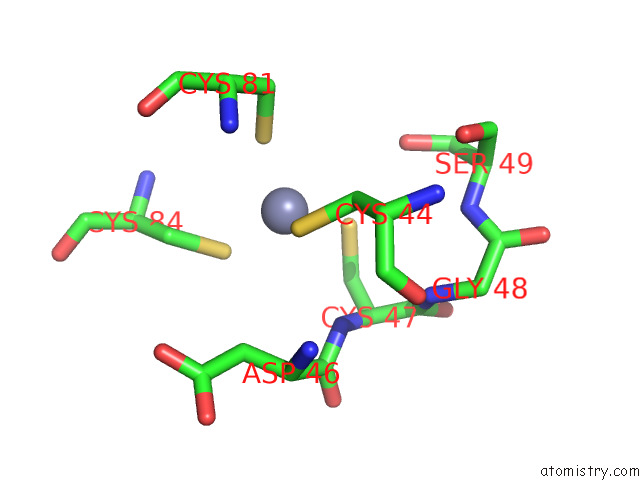
Mono view
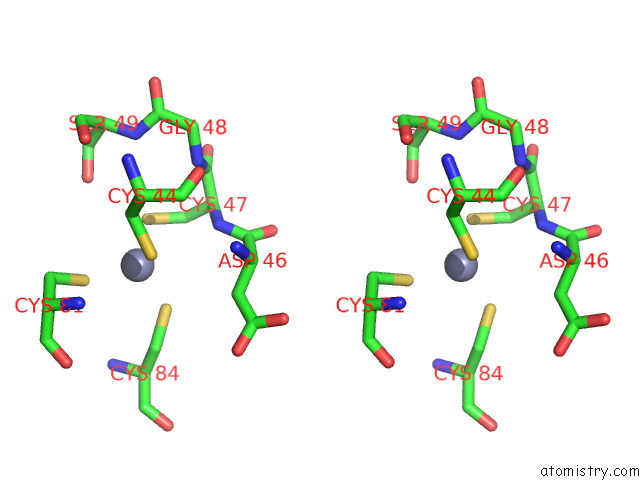
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of State E2 Nucleolar 60S Ribosomal Biogenesis Intermediate - SPB4 Local Refinement Model within 5.0Å range:
|
Reference:
V.E.Cruz,
K.Sekulski,
N.Peddada,
C.Sailer,
S.Balasubramanian,
C.S.Weirich,
F.Stengel,
J.P.Erzberger.
Sequence-Specific Remodeling of A Topologically Complex Rnp Substrate By SPB4. Nat.Struct.Mol.Biol. V. 29 1228 2022.
ISSN: ESSN 1545-9985
PubMed: 36482249
DOI: 10.1038/S41594-022-00874-9
Page generated: Wed Oct 30 07:42:46 2024
ISSN: ESSN 1545-9985
PubMed: 36482249
DOI: 10.1038/S41594-022-00874-9
Last articles
Zn in 9MJ5Zn in 9HNW
Zn in 9G0L
Zn in 9FNE
Zn in 9DZN
Zn in 9E0I
Zn in 9D32
Zn in 9DAK
Zn in 8ZXC
Zn in 8ZUF