Zinc »
PDB 3no5-3o7a »
3nqz »
Zinc in PDB 3nqz: Crystal Structure of the Autoprocessed Vibriolysin Mcp-02 with E369A Mutation
Enzymatic activity of Crystal Structure of the Autoprocessed Vibriolysin Mcp-02 with E369A Mutation
All present enzymatic activity of Crystal Structure of the Autoprocessed Vibriolysin Mcp-02 with E369A Mutation:
3.4.24.25;
3.4.24.25;
Protein crystallography data
The structure of Crystal Structure of the Autoprocessed Vibriolysin Mcp-02 with E369A Mutation, PDB code: 3nqz
was solved by
X.Gao,
J.Wang,
L.Chen,
J.-W.Wu,
Y.-Z.Zhang,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 16.97 / 2.05 |
| Space group | P 32 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 82.535, 82.535, 154.206, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 17.9 / 22 |
Other elements in 3nqz:
The structure of Crystal Structure of the Autoprocessed Vibriolysin Mcp-02 with E369A Mutation also contains other interesting chemical elements:
| Calcium | (Ca) | 1 atom |
Zinc Binding Sites:
The binding sites of Zinc atom in the Crystal Structure of the Autoprocessed Vibriolysin Mcp-02 with E369A Mutation
(pdb code 3nqz). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total only one binding site of Zinc was determined in the Crystal Structure of the Autoprocessed Vibriolysin Mcp-02 with E369A Mutation, PDB code: 3nqz:
In total only one binding site of Zinc was determined in the Crystal Structure of the Autoprocessed Vibriolysin Mcp-02 with E369A Mutation, PDB code: 3nqz:
Zinc binding site 1 out of 1 in 3nqz
Go back to
Zinc binding site 1 out
of 1 in the Crystal Structure of the Autoprocessed Vibriolysin Mcp-02 with E369A Mutation
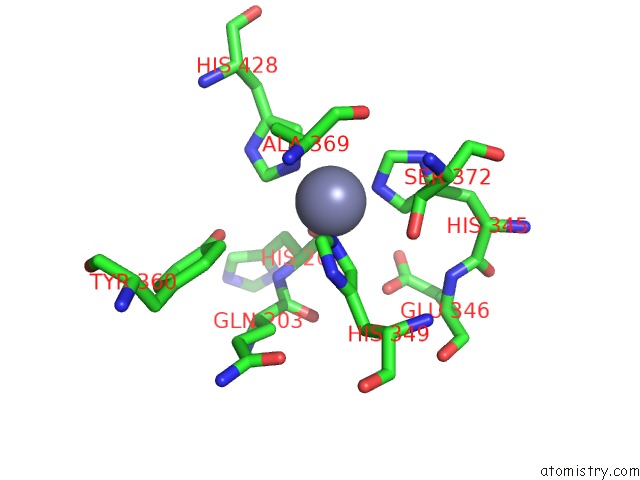
Mono view
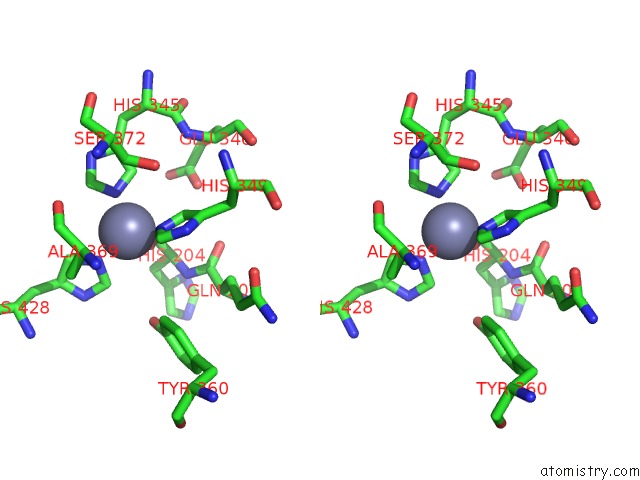
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Crystal Structure of the Autoprocessed Vibriolysin Mcp-02 with E369A Mutation within 5.0Å range:
|
Reference:
X.Gao,
J.Wang,
D.-Q.Yu,
F.Bian,
B.-B.Xie,
X.-L.Chen,
B.-C.Zhou,
L.-H.Lai,
Z.-X.Wang,
J.-W.Wu,
Y.-Z.Zhang.
Structural Basis For the Autoprocessing of Zinc Metalloproteases in the Thermolysin Family Proc.Natl.Acad.Sci.Usa V. 107 17569 2010.
ISSN: ISSN 0027-8424
PubMed: 20876133
DOI: 10.1073/PNAS.1005681107
Page generated: Wed Aug 20 12:24:33 2025
ISSN: ISSN 0027-8424
PubMed: 20876133
DOI: 10.1073/PNAS.1005681107
Last articles
Zn in 4HMHZn in 4HLM
Zn in 4HLK
Zn in 4HL2
Zn in 4HLH
Zn in 4HLG
Zn in 4HLF
Zn in 4HL5
Zn in 4HKN
Zn in 4HKR