Zinc »
PDB 2yqq-2yu4 »
2ysm »
Zinc in PDB 2ysm: Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3)
Enzymatic activity of Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3)
All present enzymatic activity of Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3):
2.1.1.43;
2.1.1.43;
Zinc Binding Sites:
The binding sites of Zinc atom in the Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3)
(pdb code 2ysm). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total 4 binding sites of Zinc where determined in the Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3), PDB code: 2ysm:
Jump to Zinc binding site number: 1; 2; 3; 4;
In total 4 binding sites of Zinc where determined in the Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3), PDB code: 2ysm:
Jump to Zinc binding site number: 1; 2; 3; 4;
Zinc binding site 1 out of 4 in 2ysm
Go back to
Zinc binding site 1 out
of 4 in the Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3)
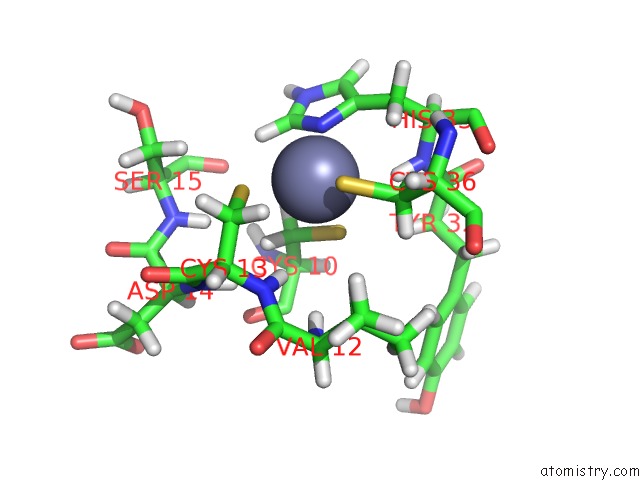
Mono view
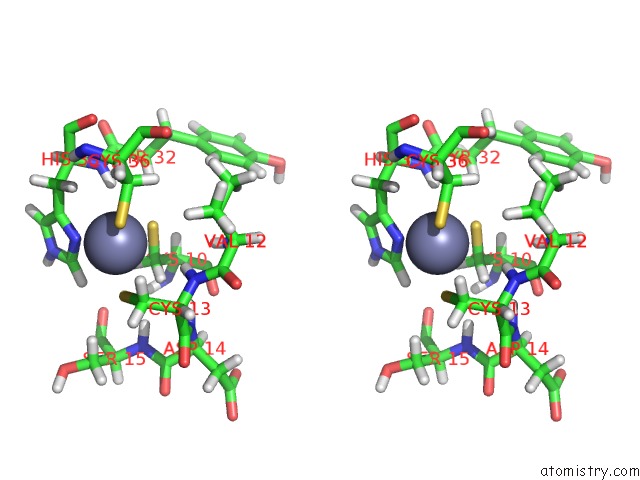
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3) within 5.0Å range:
|
Zinc binding site 2 out of 4 in 2ysm
Go back to
Zinc binding site 2 out
of 4 in the Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3)
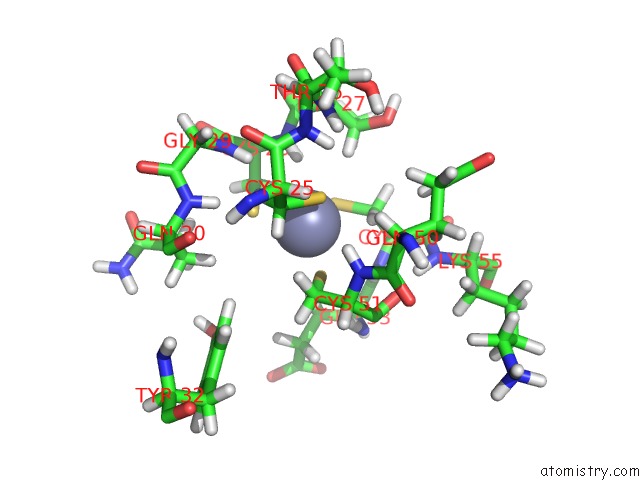
Mono view
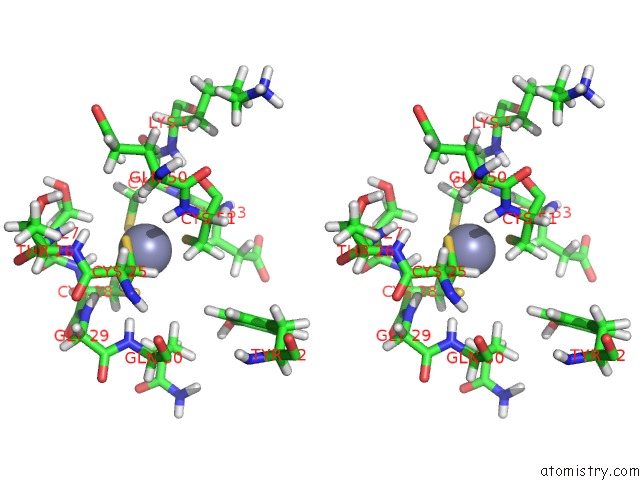
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 2 of Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3) within 5.0Å range:
|
Zinc binding site 3 out of 4 in 2ysm
Go back to
Zinc binding site 3 out
of 4 in the Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3)
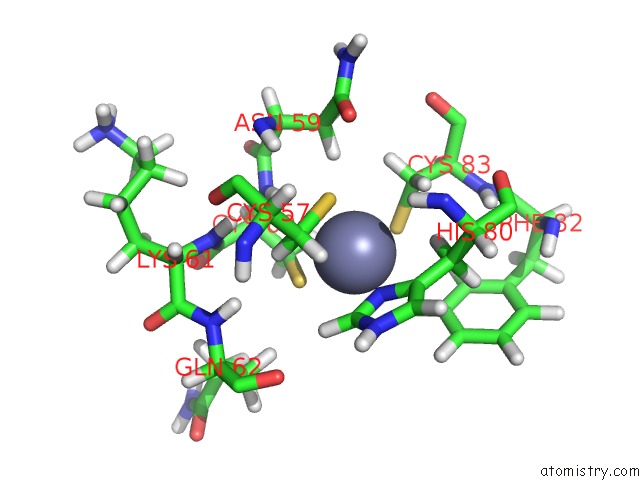
Mono view
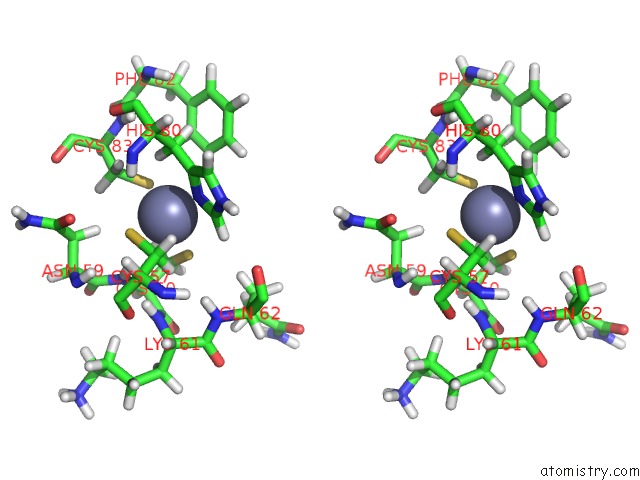
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 3 of Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3) within 5.0Å range:
|
Zinc binding site 4 out of 4 in 2ysm
Go back to
Zinc binding site 4 out
of 4 in the Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3)
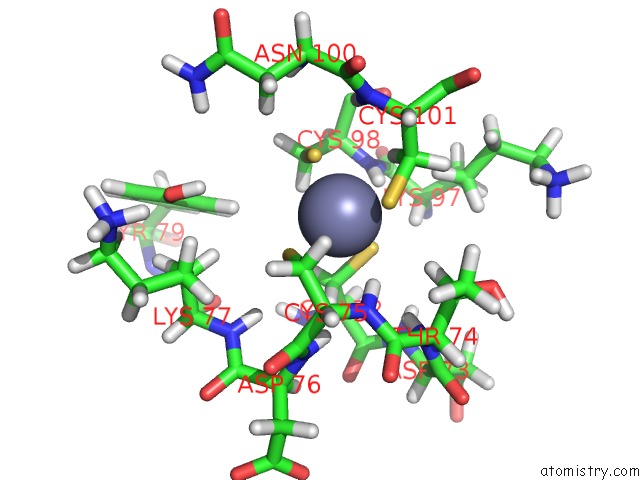
Mono view
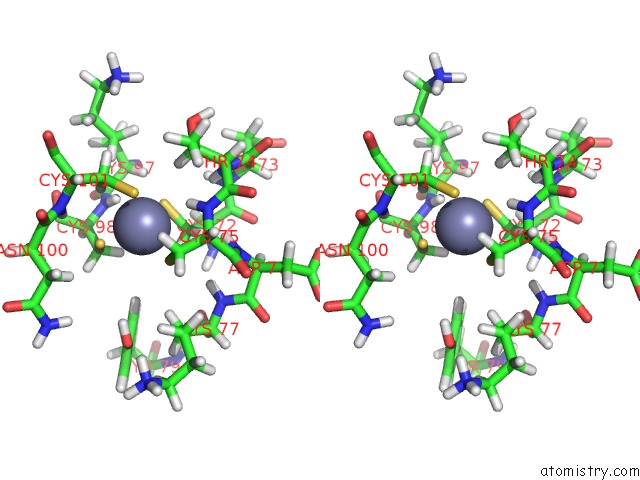
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 4 of Solution Structure of the Second and Third Phd Domain From Histone- Lysine N-Methyltransferase 2C (KMT2C/MLL3) within 5.0Å range:
|
Reference:
X.R.Qin,
F.Hayashi,
S.Yokoyama.
Solution Structure of the First and Second Phd Domain From Myeloid/Lymphoid or Mixed-Lineage Leukemia Protein 3 Homolog To Be Published.
Page generated: Thu Oct 24 10:23:52 2024
Last articles
Zn in 9J0NZn in 9J0O
Zn in 9J0P
Zn in 9FJX
Zn in 9EKB
Zn in 9C0F
Zn in 9CAH
Zn in 9CH0
Zn in 9CH3
Zn in 9CH1