Zinc »
PDB 2hqk-2ibi »
2i50 »
Zinc in PDB 2i50: Solution Structure of Ubp-M Znf-Ubp Domain
Enzymatic activity of Solution Structure of Ubp-M Znf-Ubp Domain
All present enzymatic activity of Solution Structure of Ubp-M Znf-Ubp Domain:
3.1.2.15;
3.1.2.15;
Zinc Binding Sites:
The binding sites of Zinc atom in the Solution Structure of Ubp-M Znf-Ubp Domain
(pdb code 2i50). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total 3 binding sites of Zinc where determined in the Solution Structure of Ubp-M Znf-Ubp Domain, PDB code: 2i50:
Jump to Zinc binding site number: 1; 2; 3;
In total 3 binding sites of Zinc where determined in the Solution Structure of Ubp-M Znf-Ubp Domain, PDB code: 2i50:
Jump to Zinc binding site number: 1; 2; 3;
Zinc binding site 1 out of 3 in 2i50
Go back to
Zinc binding site 1 out
of 3 in the Solution Structure of Ubp-M Znf-Ubp Domain
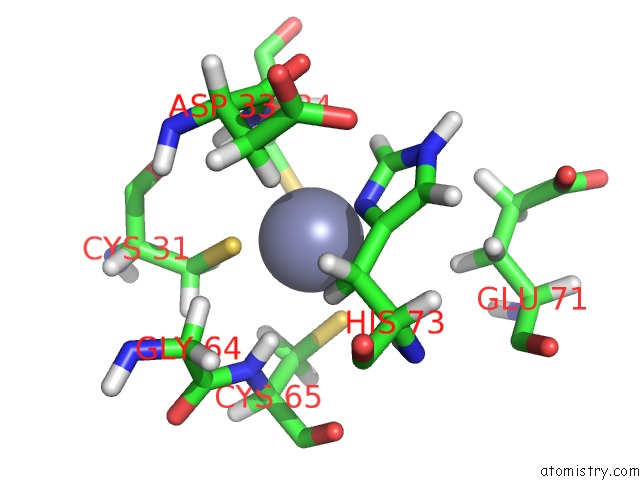
Mono view
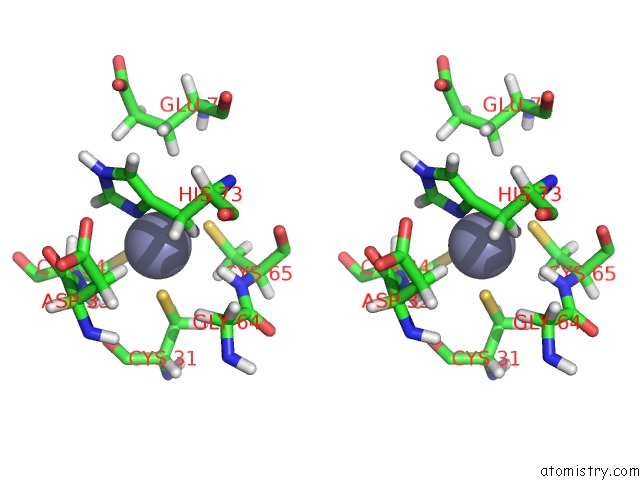
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Solution Structure of Ubp-M Znf-Ubp Domain within 5.0Å range:
|
Zinc binding site 2 out of 3 in 2i50
Go back to
Zinc binding site 2 out
of 3 in the Solution Structure of Ubp-M Znf-Ubp Domain
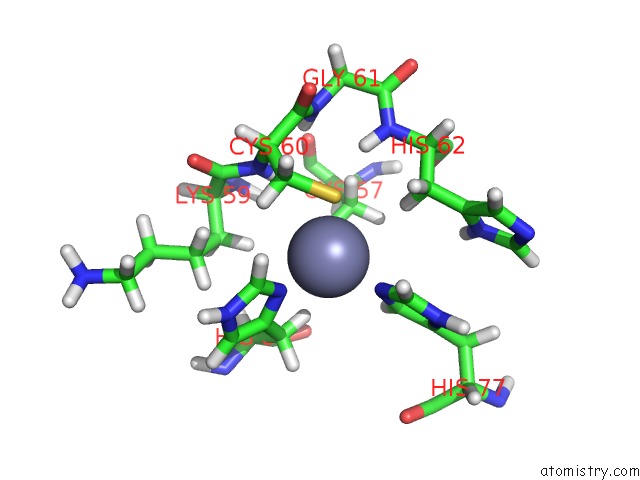
Mono view
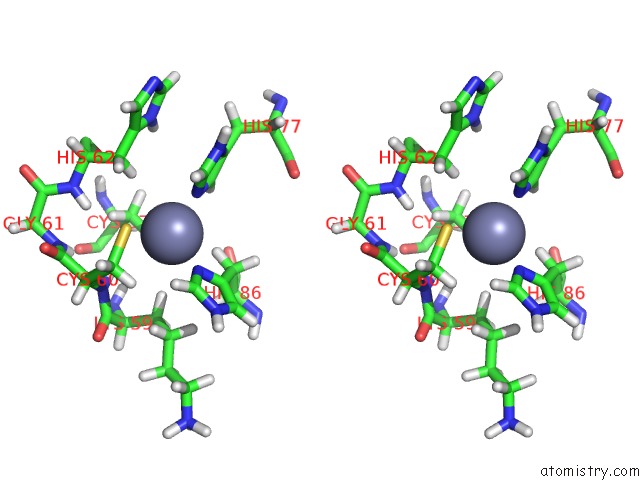
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 2 of Solution Structure of Ubp-M Znf-Ubp Domain within 5.0Å range:
|
Zinc binding site 3 out of 3 in 2i50
Go back to
Zinc binding site 3 out
of 3 in the Solution Structure of Ubp-M Znf-Ubp Domain
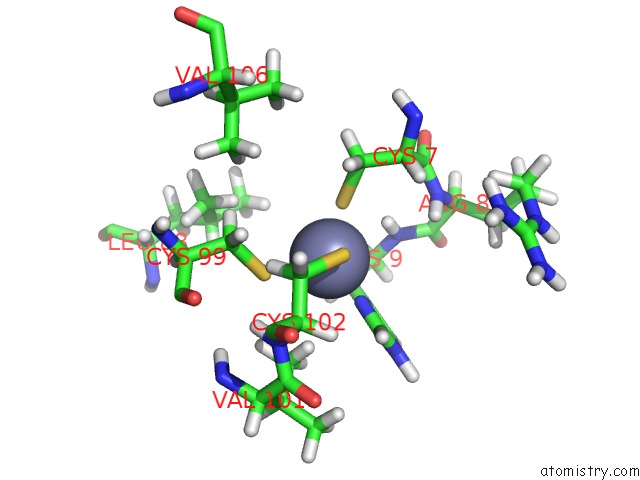
Mono view
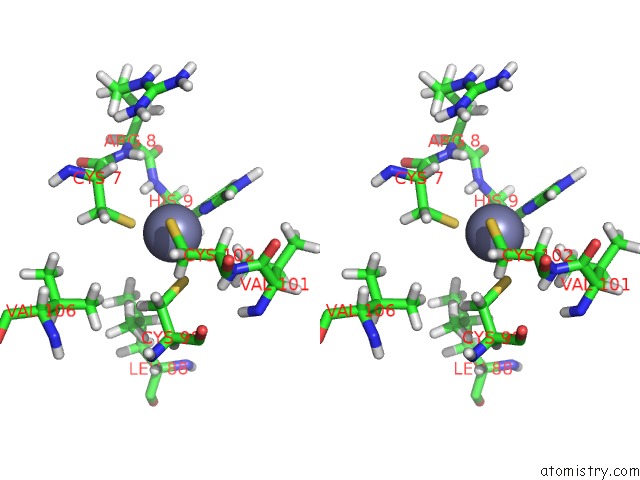
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 3 of Solution Structure of Ubp-M Znf-Ubp Domain within 5.0Å range:
|
Reference:
M.T.Pai,
S.R.Tzeng,
J.J.Kovacs,
M.A.Keaton,
S.S.Li,
T.P.Yao,
P.Zhou.
Solution Structure of the Ubp-M Buz Domain, A Highly Specific Protein Module That Recognizes the C-Terminal Tail of Free Ubiquitin. J.Mol.Biol. V. 370 290 2007.
ISSN: ISSN 0022-2836
PubMed: 17512543
DOI: 10.1016/J.JMB.2007.04.015
Page generated: Thu Oct 17 00:50:26 2024
ISSN: ISSN 0022-2836
PubMed: 17512543
DOI: 10.1016/J.JMB.2007.04.015
Last articles
Zn in 9J0NZn in 9J0O
Zn in 9J0P
Zn in 9FJX
Zn in 9EKB
Zn in 9C0F
Zn in 9CAH
Zn in 9CH0
Zn in 9CH3
Zn in 9CH1