Zinc »
PDB 1sdz-1sxb »
1sp2 »
Zinc in PDB 1sp2: uc(Nmr) Structure of A Zinc Finger Domain From Transcription Factor SP1F2, Minimized Average Structure
Zinc Binding Sites:
The binding sites of Zinc atom in the uc(Nmr) Structure of A Zinc Finger Domain From Transcription Factor SP1F2, Minimized Average Structure
(pdb code 1sp2). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total only one binding site of Zinc was determined in the uc(Nmr) Structure of A Zinc Finger Domain From Transcription Factor SP1F2, Minimized Average Structure, PDB code: 1sp2:
In total only one binding site of Zinc was determined in the uc(Nmr) Structure of A Zinc Finger Domain From Transcription Factor SP1F2, Minimized Average Structure, PDB code: 1sp2:
Zinc binding site 1 out of 1 in 1sp2
Go back to
Zinc binding site 1 out
of 1 in the uc(Nmr) Structure of A Zinc Finger Domain From Transcription Factor SP1F2, Minimized Average Structure
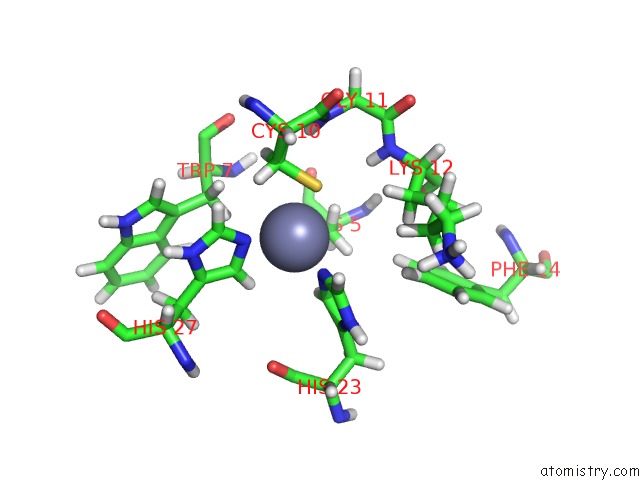
Mono view
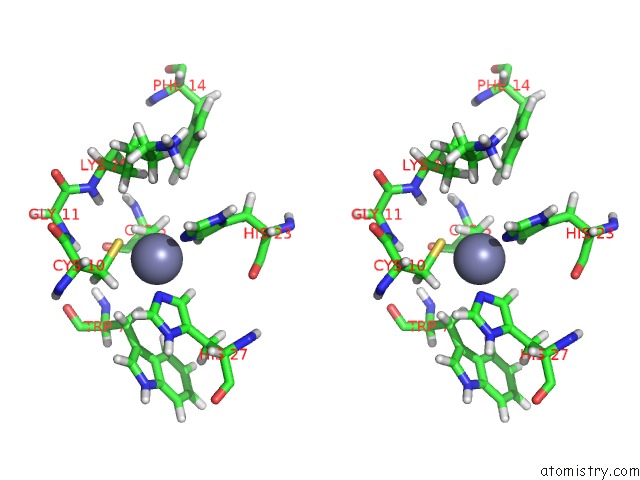
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of uc(Nmr) Structure of A Zinc Finger Domain From Transcription Factor SP1F2, Minimized Average Structure within 5.0Å range:
|
Reference:
V.A.Narayan,
R.W.Kriwacki,
J.P.Caradonna.
Structures of Zinc Finger Domains From Transcription Factor SP1. Insights Into Sequence-Specific Protein-Dna Recognition. J.Biol.Chem. V. 272 7801 1997.
ISSN: ISSN 0021-9258
PubMed: 9065444
DOI: 10.1074/JBC.272.12.7801
Page generated: Wed Oct 16 18:53:41 2024
ISSN: ISSN 0021-9258
PubMed: 9065444
DOI: 10.1074/JBC.272.12.7801
Last articles
Zn in 9J0NZn in 9J0O
Zn in 9J0P
Zn in 9FJX
Zn in 9EKB
Zn in 9C0F
Zn in 9CAH
Zn in 9CH0
Zn in 9CH3
Zn in 9CH1