Zinc »
PDB 1oj7-1p1r »
1p0c »
Zinc in PDB 1p0c: Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8)
Enzymatic activity of Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8)
All present enzymatic activity of Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8):
1.1.1.2;
1.1.1.2;
Protein crystallography data
The structure of Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8), PDB code: 1p0c
was solved by
A.Rosell,
E.Valencia,
X.Pares,
I.Fita,
J.Farres,
W.F.Ochoa,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 30.00 / 2.20 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 122.551, 79.523, 91.892, 90.00, 113.10, 90.00 |
| R / Rfree (%) | 19.8 / 23.4 |
Zinc Binding Sites:
The binding sites of Zinc atom in the Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8)
(pdb code 1p0c). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total 4 binding sites of Zinc where determined in the Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8), PDB code: 1p0c:
Jump to Zinc binding site number: 1; 2; 3; 4;
In total 4 binding sites of Zinc where determined in the Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8), PDB code: 1p0c:
Jump to Zinc binding site number: 1; 2; 3; 4;
Zinc binding site 1 out of 4 in 1p0c
Go back to
Zinc binding site 1 out
of 4 in the Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8)
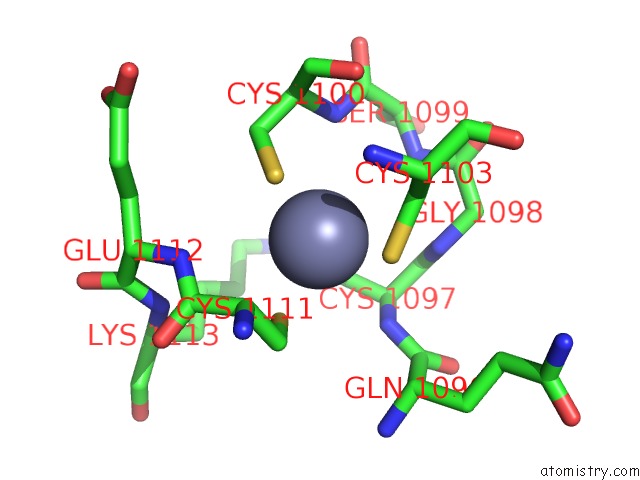
Mono view
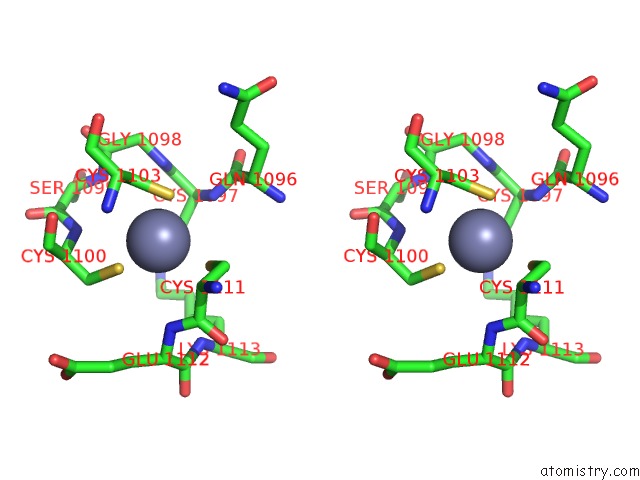
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8) within 5.0Å range:
|
Zinc binding site 2 out of 4 in 1p0c
Go back to
Zinc binding site 2 out
of 4 in the Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8)
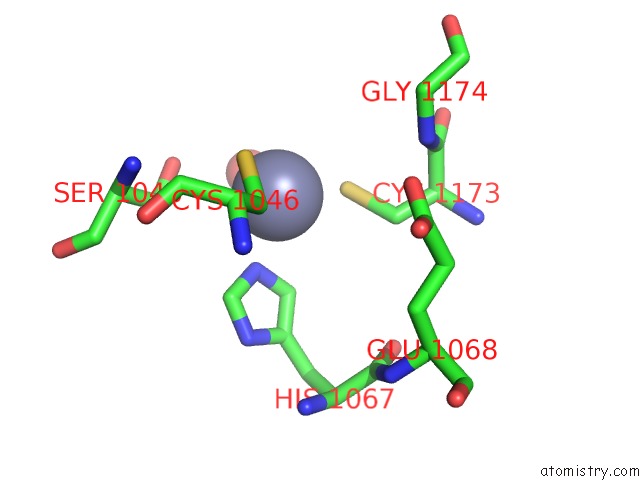
Mono view
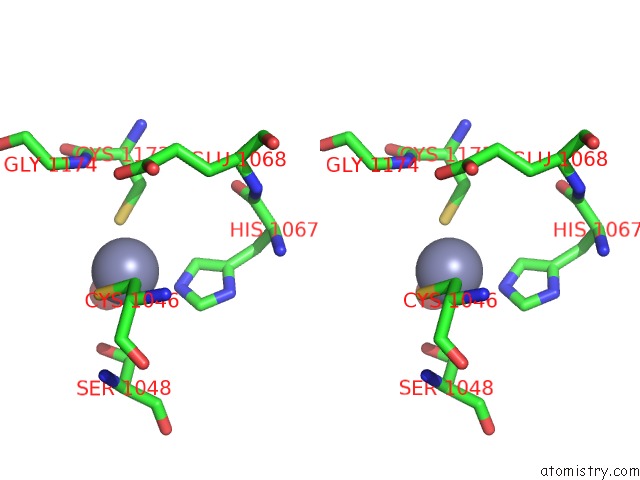
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 2 of Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8) within 5.0Å range:
|
Zinc binding site 3 out of 4 in 1p0c
Go back to
Zinc binding site 3 out
of 4 in the Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8)
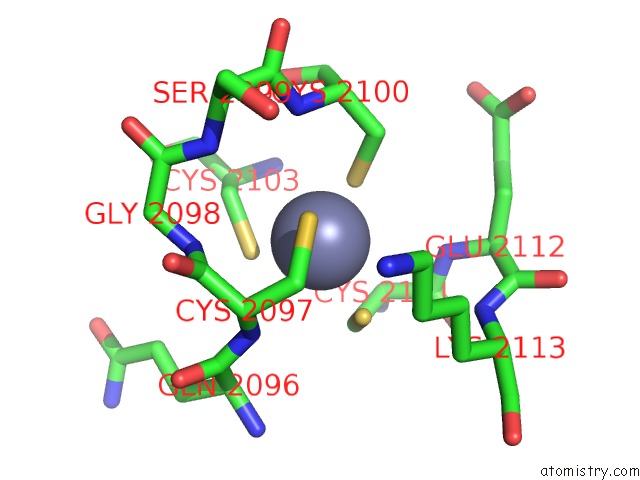
Mono view
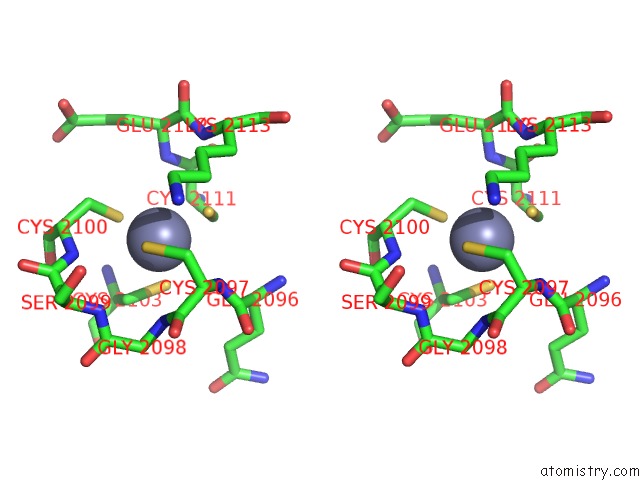
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 3 of Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8) within 5.0Å range:
|
Zinc binding site 4 out of 4 in 1p0c
Go back to
Zinc binding site 4 out
of 4 in the Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8)
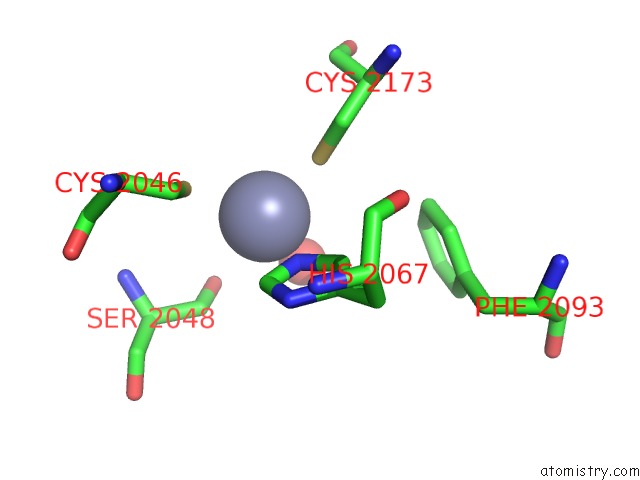
Mono view
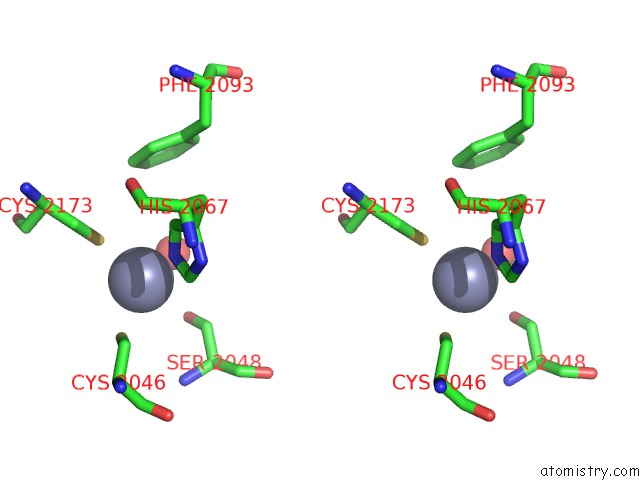
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 4 of Crystal Structure of the Nadp(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8) within 5.0Å range:
|
Reference:
A.Rosell,
E.Valencia,
X.Pares,
I.Fita,
J.Farres,
W.F.Ochoa.
Crystal Structure of the Vertebrate Nadp(H)-Dependent Alcohol Dehydrogenase (ADH8) J.Mol.Biol. V. 330 75 2003.
ISSN: ISSN 0022-2836
PubMed: 12818203
DOI: 10.1016/S0022-2836(03)00431-5
Page generated: Wed Oct 16 17:39:13 2024
ISSN: ISSN 0022-2836
PubMed: 12818203
DOI: 10.1016/S0022-2836(03)00431-5
Last articles
Zn in 9J0NZn in 9J0O
Zn in 9J0P
Zn in 9FJX
Zn in 9EKB
Zn in 9C0F
Zn in 9CAH
Zn in 9CH0
Zn in 9CH3
Zn in 9CH1