Zinc »
PDB 6gco-6gnq »
6gkh »
Zinc in PDB 6gkh: Cryoem Structure of the MDA5-Dsrna Filament in Complex with Adp-ALF4
Enzymatic activity of Cryoem Structure of the MDA5-Dsrna Filament in Complex with Adp-ALF4
All present enzymatic activity of Cryoem Structure of the MDA5-Dsrna Filament in Complex with Adp-ALF4:
3.6.4.13;
3.6.4.13;
Other elements in 6gkh:
The structure of Cryoem Structure of the MDA5-Dsrna Filament in Complex with Adp-ALF4 also contains other interesting chemical elements:
| Fluorine | (F) | 4 atoms |
| Magnesium | (Mg) | 1 atom |
| Aluminium | (Al) | 1 atom |
Zinc Binding Sites:
The binding sites of Zinc atom in the Cryoem Structure of the MDA5-Dsrna Filament in Complex with Adp-ALF4
(pdb code 6gkh). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total only one binding site of Zinc was determined in the Cryoem Structure of the MDA5-Dsrna Filament in Complex with Adp-ALF4, PDB code: 6gkh:
In total only one binding site of Zinc was determined in the Cryoem Structure of the MDA5-Dsrna Filament in Complex with Adp-ALF4, PDB code: 6gkh:
Zinc binding site 1 out of 1 in 6gkh
Go back to
Zinc binding site 1 out
of 1 in the Cryoem Structure of the MDA5-Dsrna Filament in Complex with Adp-ALF4
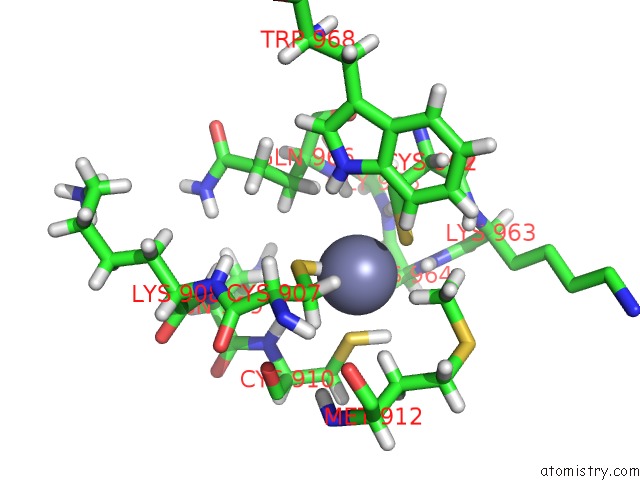
Mono view
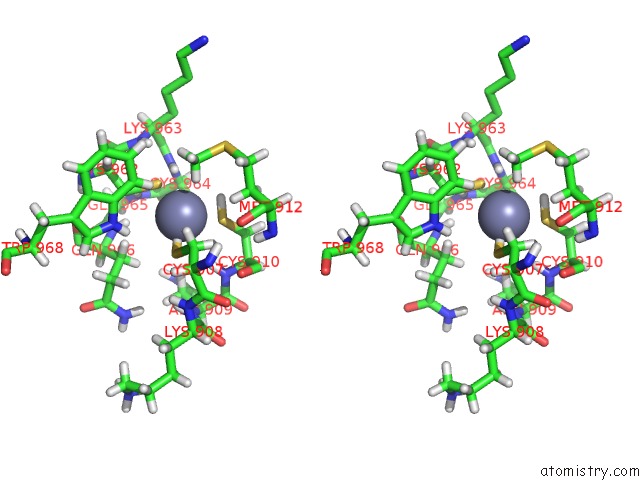
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Cryoem Structure of the MDA5-Dsrna Filament in Complex with Adp-ALF4 within 5.0Å range:
|
Reference:
Q.Yu,
K.Qu,
Y.Modis.
Cryo-Em Structures of MDA5-Dsrna Filaments at Different Stages of Atp Hydrolysis. Mol. Cell V. 72 999 2018.
ISSN: ISSN 1097-4164
PubMed: 30449722
DOI: 10.1016/J.MOLCEL.2018.10.012
Page generated: Mon Oct 28 21:53:33 2024
ISSN: ISSN 1097-4164
PubMed: 30449722
DOI: 10.1016/J.MOLCEL.2018.10.012
Last articles
Zn in 9MJ5Zn in 9HNW
Zn in 9G0L
Zn in 9FNE
Zn in 9DZN
Zn in 9E0I
Zn in 9D32
Zn in 9DAK
Zn in 8ZXC
Zn in 8ZUF