Zinc »
PDB 5a25-5aba »
5a50 »
Zinc in PDB 5a50: The Crystal Structure of Arabidopsis Thaliana CAR4 in Complex with Two Calcium Ions, Zn and Phopho Choline
Protein crystallography data
The structure of The Crystal Structure of Arabidopsis Thaliana CAR4 in Complex with Two Calcium Ions, Zn and Phopho Choline, PDB code: 5a50
was solved by
M.Diaz,
A.Albert,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 47.56 / 2.40 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 35.737, 89.150, 112.475, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19.5 / 24.5 |
Other elements in 5a50:
The structure of The Crystal Structure of Arabidopsis Thaliana CAR4 in Complex with Two Calcium Ions, Zn and Phopho Choline also contains other interesting chemical elements:
| Calcium | (Ca) | 4 atoms |
Zinc Binding Sites:
The binding sites of Zinc atom in the The Crystal Structure of Arabidopsis Thaliana CAR4 in Complex with Two Calcium Ions, Zn and Phopho Choline
(pdb code 5a50). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total only one binding site of Zinc was determined in the The Crystal Structure of Arabidopsis Thaliana CAR4 in Complex with Two Calcium Ions, Zn and Phopho Choline, PDB code: 5a50:
In total only one binding site of Zinc was determined in the The Crystal Structure of Arabidopsis Thaliana CAR4 in Complex with Two Calcium Ions, Zn and Phopho Choline, PDB code: 5a50:
Zinc binding site 1 out of 1 in 5a50
Go back to
Zinc binding site 1 out
of 1 in the The Crystal Structure of Arabidopsis Thaliana CAR4 in Complex with Two Calcium Ions, Zn and Phopho Choline
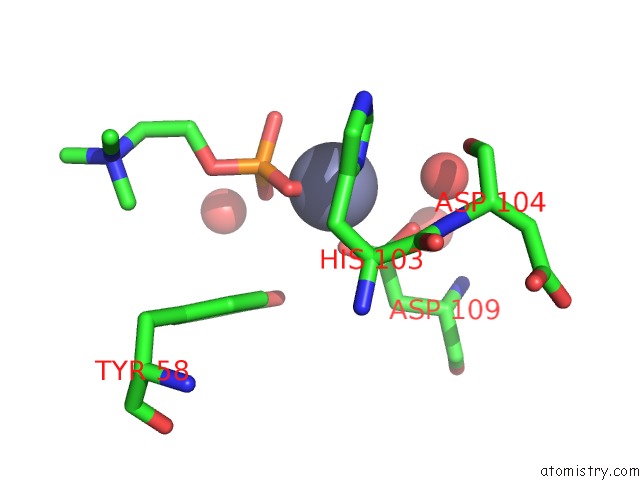
Mono view
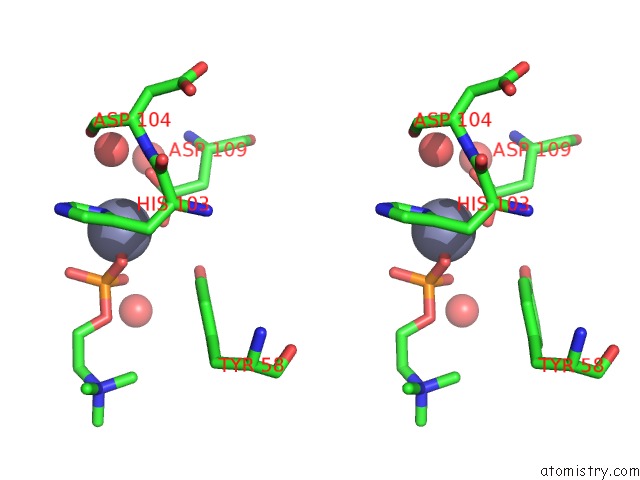
Stereo pair view

Mono view
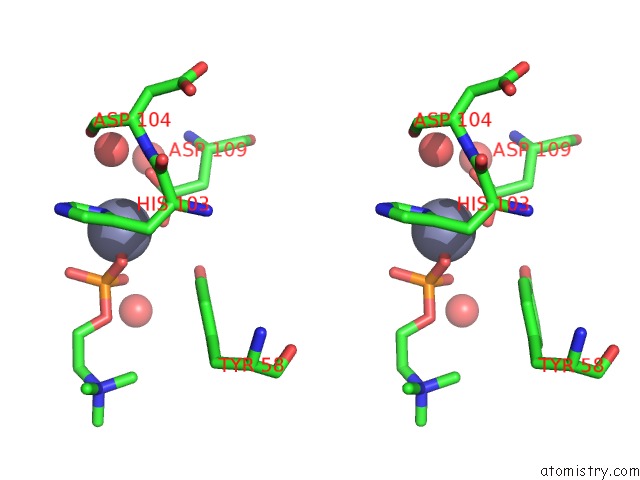
Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of The Crystal Structure of Arabidopsis Thaliana CAR4 in Complex with Two Calcium Ions, Zn and Phopho Choline within 5.0Å range:
|
Reference:
M.Diaz,
M.J.Sanchez-Barrena,
J.M.Gonzalez-Rubio,
L.Rodriguez,
D.Fernandez,
R.Antoni,
C.Yunta,
B.Belda-Palazon,
M.Gonzalez-Guzman,
M.Peirats-Llobet,
M.Menendez,
J.Boskovic,
J.A.Marquez,
P.L.Rodriguez,
A.Albert.
Calcium-Dependent Oligomerization of Car Proteins at Cell Membrane Modulates Aba Signaling. Proc.Natl.Acad.Sci.Usa V. 113 E396 2016.
ISSN: ISSN 0027-8424
PubMed: 26719420
DOI: 10.1073/PNAS.1512779113
Page generated: Thu Aug 21 00:32:31 2025
ISSN: ISSN 0027-8424
PubMed: 26719420
DOI: 10.1073/PNAS.1512779113
Last articles
Zn in 5RLHZn in 5RLG
Zn in 5RLF
Zn in 5RLE
Zn in 5RLD
Zn in 5RLB
Zn in 5RLC
Zn in 5RL9
Zn in 5RL8
Zn in 5RL6