Zinc »
PDB 4g3m-4ggj »
4gat »
Zinc in PDB 4gat: Solution uc(Nmr) Structure of the Wild Type Dna Binding Domain of Area Complexed to A 13BP Dna Containing A Cgata Site, Regularized Mean Structure
Zinc Binding Sites:
The binding sites of Zinc atom in the Solution uc(Nmr) Structure of the Wild Type Dna Binding Domain of Area Complexed to A 13BP Dna Containing A Cgata Site, Regularized Mean Structure
(pdb code 4gat). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total only one binding site of Zinc was determined in the Solution uc(Nmr) Structure of the Wild Type Dna Binding Domain of Area Complexed to A 13BP Dna Containing A Cgata Site, Regularized Mean Structure, PDB code: 4gat:
In total only one binding site of Zinc was determined in the Solution uc(Nmr) Structure of the Wild Type Dna Binding Domain of Area Complexed to A 13BP Dna Containing A Cgata Site, Regularized Mean Structure, PDB code: 4gat:
Zinc binding site 1 out of 1 in 4gat
Go back to
Zinc binding site 1 out
of 1 in the Solution uc(Nmr) Structure of the Wild Type Dna Binding Domain of Area Complexed to A 13BP Dna Containing A Cgata Site, Regularized Mean Structure
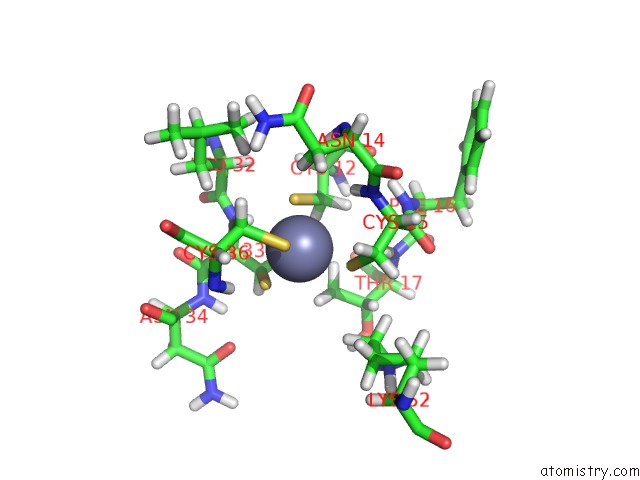
Mono view
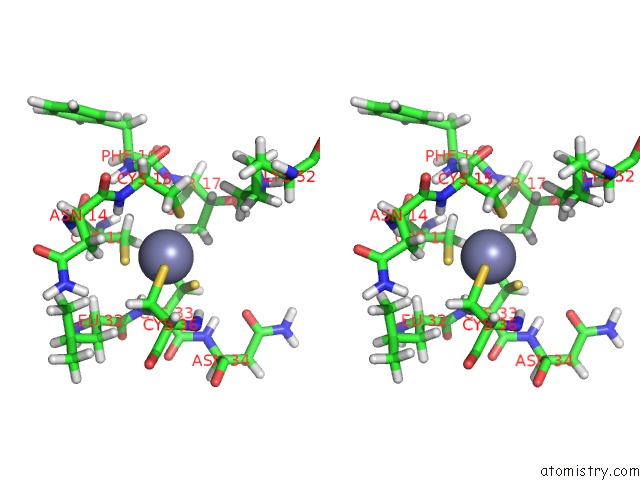
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Solution uc(Nmr) Structure of the Wild Type Dna Binding Domain of Area Complexed to A 13BP Dna Containing A Cgata Site, Regularized Mean Structure within 5.0Å range:
|
Reference:
M.R.Starich,
M.Wikstrom,
H.N.Arst Jr.,
G.M.Clore,
A.M.Gronenborn.
The Solution Structure of A Fungal Area Protein-Dna Complex: An Alternative Binding Mode For the Basic Carboxyl Tail of Gata Factors. J.Mol.Biol. V. 277 605 1998.
ISSN: ISSN 0022-2836
PubMed: 9533883
DOI: 10.1006/JMBI.1998.1625
Page generated: Sat Oct 26 23:12:30 2024
ISSN: ISSN 0022-2836
PubMed: 9533883
DOI: 10.1006/JMBI.1998.1625
Last articles
Zn in 9MJ5Zn in 9HNW
Zn in 9G0L
Zn in 9FNE
Zn in 9DZN
Zn in 9E0I
Zn in 9D32
Zn in 9DAK
Zn in 8ZXC
Zn in 8ZUF