Zinc »
PDB 4fkh-4fvu »
4fup »
Zinc in PDB 4fup: Structural Basis For ZN2+-Dependent Intercellular Adhesion in Staphylococcal Biofilms
Protein crystallography data
The structure of Structural Basis For ZN2+-Dependent Intercellular Adhesion in Staphylococcal Biofilms, PDB code: 4fup
was solved by
D.G.Conrady,
J.J.Wilson,
A.B.Herr,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 45.36 / 2.51 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 83.101, 34.240, 138.779, 90.00, 101.31, 90.00 |
| R / Rfree (%) | 21.9 / 27.6 |
Zinc Binding Sites:
The binding sites of Zinc atom in the Structural Basis For ZN2+-Dependent Intercellular Adhesion in Staphylococcal Biofilms
(pdb code 4fup). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total 2 binding sites of Zinc where determined in the Structural Basis For ZN2+-Dependent Intercellular Adhesion in Staphylococcal Biofilms, PDB code: 4fup:
Jump to Zinc binding site number: 1; 2;
In total 2 binding sites of Zinc where determined in the Structural Basis For ZN2+-Dependent Intercellular Adhesion in Staphylococcal Biofilms, PDB code: 4fup:
Jump to Zinc binding site number: 1; 2;
Zinc binding site 1 out of 2 in 4fup
Go back to
Zinc binding site 1 out
of 2 in the Structural Basis For ZN2+-Dependent Intercellular Adhesion in Staphylococcal Biofilms
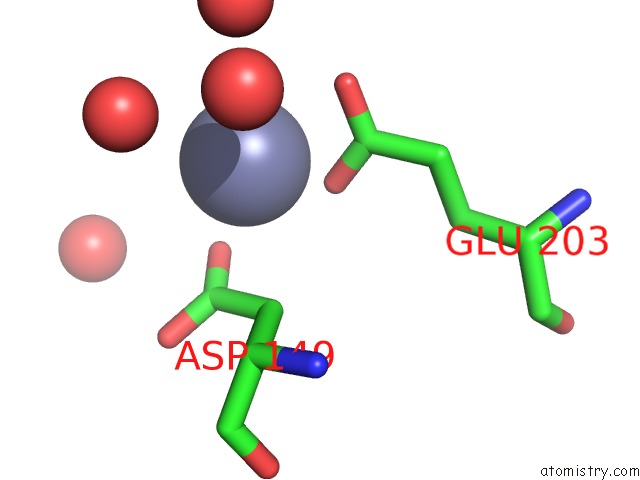
Mono view
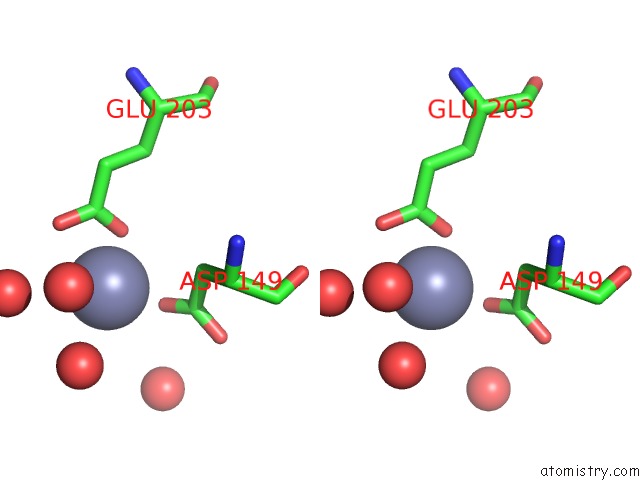
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Structural Basis For ZN2+-Dependent Intercellular Adhesion in Staphylococcal Biofilms within 5.0Å range:
|
Zinc binding site 2 out of 2 in 4fup
Go back to
Zinc binding site 2 out
of 2 in the Structural Basis For ZN2+-Dependent Intercellular Adhesion in Staphylococcal Biofilms
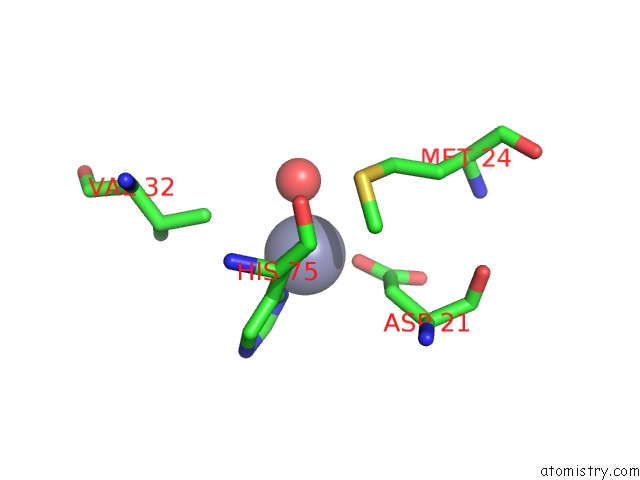
Mono view
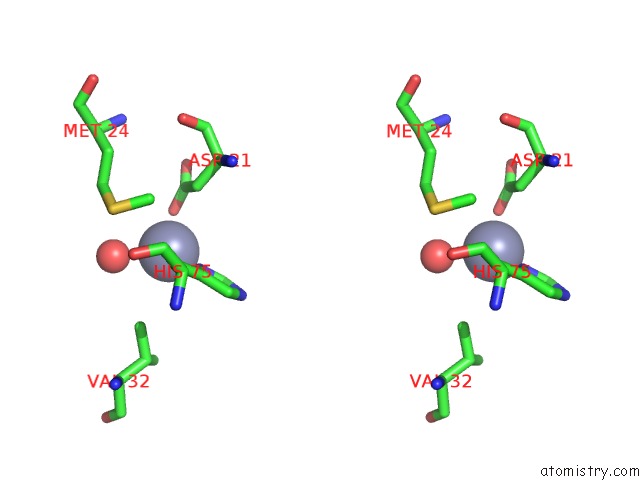
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 2 of Structural Basis For ZN2+-Dependent Intercellular Adhesion in Staphylococcal Biofilms within 5.0Å range:
|
Reference:
D.G.Conrady,
J.J.Wilson,
A.B.Herr.
Structural Basis For ZN2+-Dependent Intercellular Adhesion in Staphylococcal Biofilms. Proc.Natl.Acad.Sci.Usa V. 110 E202 2013.
ISSN: ISSN 0027-8424
PubMed: 23277549
DOI: 10.1073/PNAS.1208134110
Page generated: Sat Oct 26 22:55:04 2024
ISSN: ISSN 0027-8424
PubMed: 23277549
DOI: 10.1073/PNAS.1208134110
Last articles
Al in 4IDOAl in 4HQJ
Al in 4HNA
Al in 4GNK
Al in 4FME
Al in 4FMD
Al in 4FMC
Al in 4FMB
Al in 4C7O
Al in 4EKC