Zinc »
PDB 2vr7-2w22 »
2vrt »
Zinc in PDB 2vrt: Crystal Structure of E. Coli Rnase E Possessing M1 Rna Fragments - Catalytic Domain
Protein crystallography data
The structure of Crystal Structure of E. Coli Rnase E Possessing M1 Rna Fragments - Catalytic Domain, PDB code: 2vrt
was solved by
D.J.Koslover,
A.J.Callaghan,
M.J.Marcaida,
E.F.Garman,
M.Martick,
W.G.Scott,
B.F.Luisi,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 24.98 / 3.5 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 93.894, 121.252, 242.228, 90.00, 99.47, 90.00 |
| R / Rfree (%) | 31.7 / 35.1 |
Zinc Binding Sites:
The binding sites of Zinc atom in the Crystal Structure of E. Coli Rnase E Possessing M1 Rna Fragments - Catalytic Domain
(pdb code 2vrt). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total 2 binding sites of Zinc where determined in the Crystal Structure of E. Coli Rnase E Possessing M1 Rna Fragments - Catalytic Domain, PDB code: 2vrt:
Jump to Zinc binding site number: 1; 2;
In total 2 binding sites of Zinc where determined in the Crystal Structure of E. Coli Rnase E Possessing M1 Rna Fragments - Catalytic Domain, PDB code: 2vrt:
Jump to Zinc binding site number: 1; 2;
Zinc binding site 1 out of 2 in 2vrt
Go back to
Zinc binding site 1 out
of 2 in the Crystal Structure of E. Coli Rnase E Possessing M1 Rna Fragments - Catalytic Domain
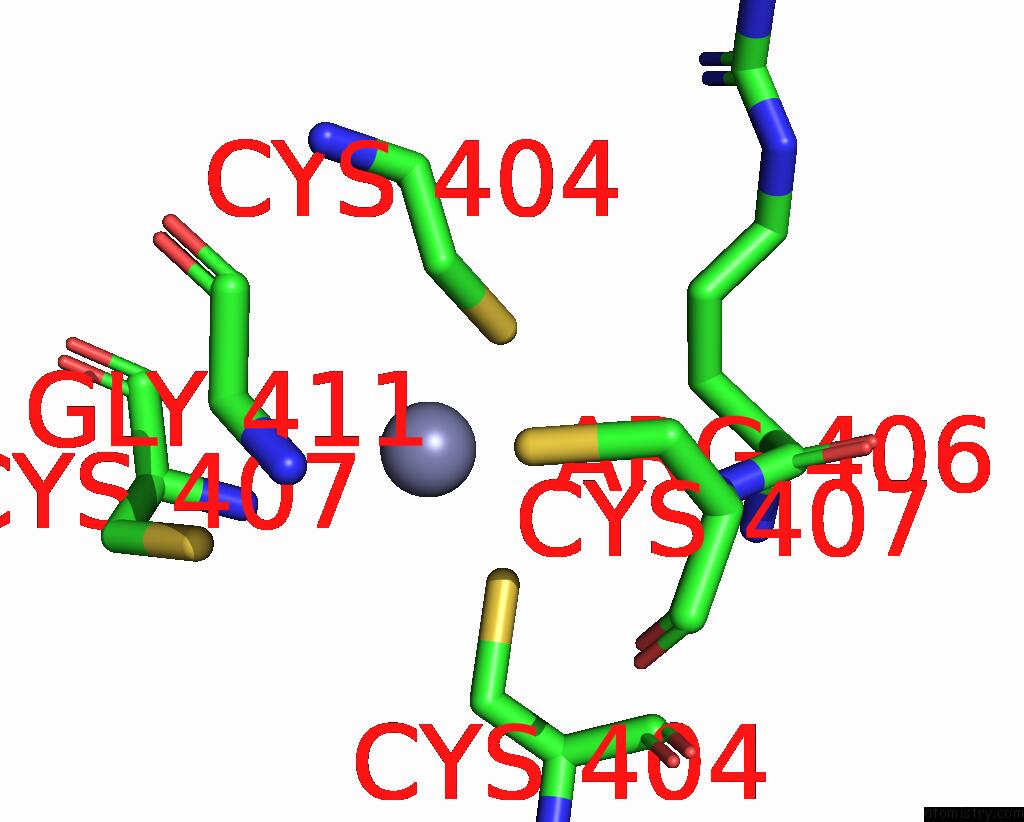
Mono view
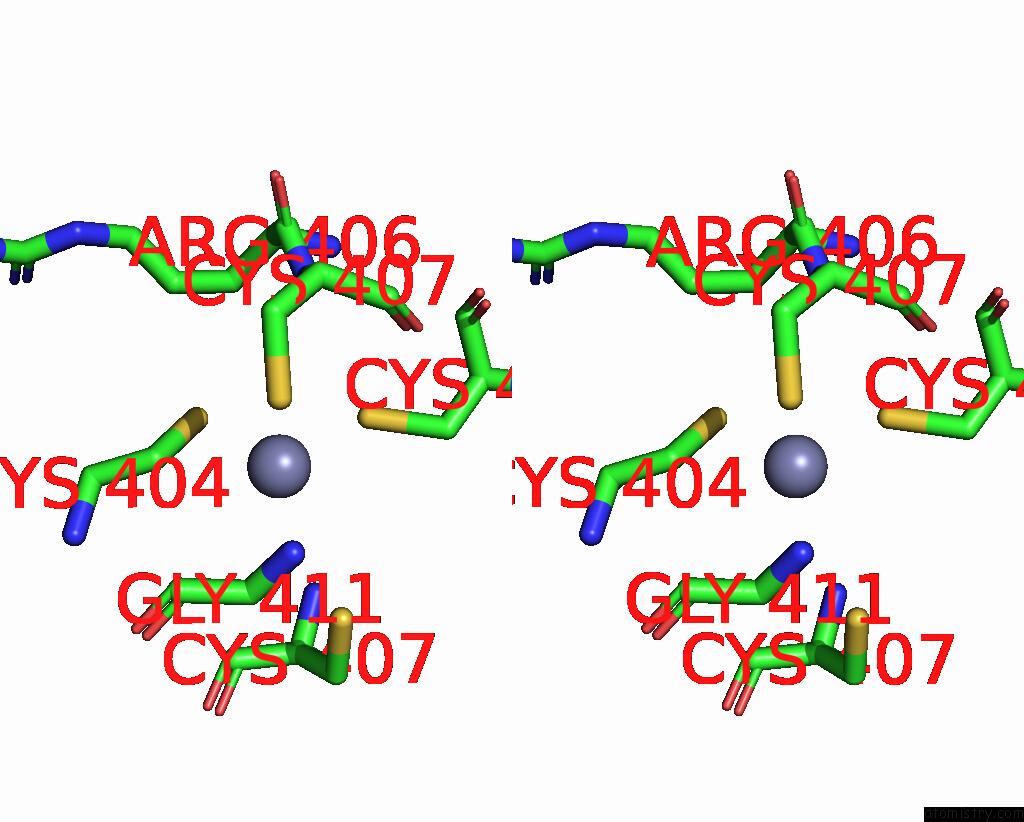
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Crystal Structure of E. Coli Rnase E Possessing M1 Rna Fragments - Catalytic Domain within 5.0Å range:
|
Zinc binding site 2 out of 2 in 2vrt
Go back to
Zinc binding site 2 out
of 2 in the Crystal Structure of E. Coli Rnase E Possessing M1 Rna Fragments - Catalytic Domain
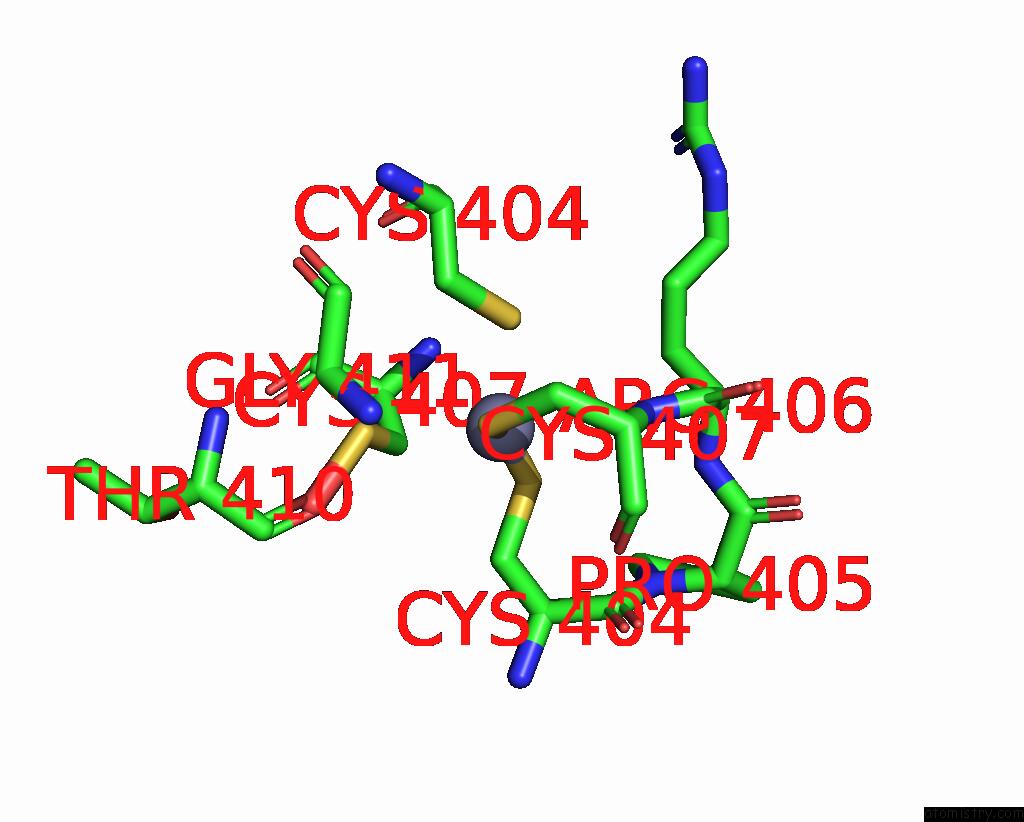
Mono view
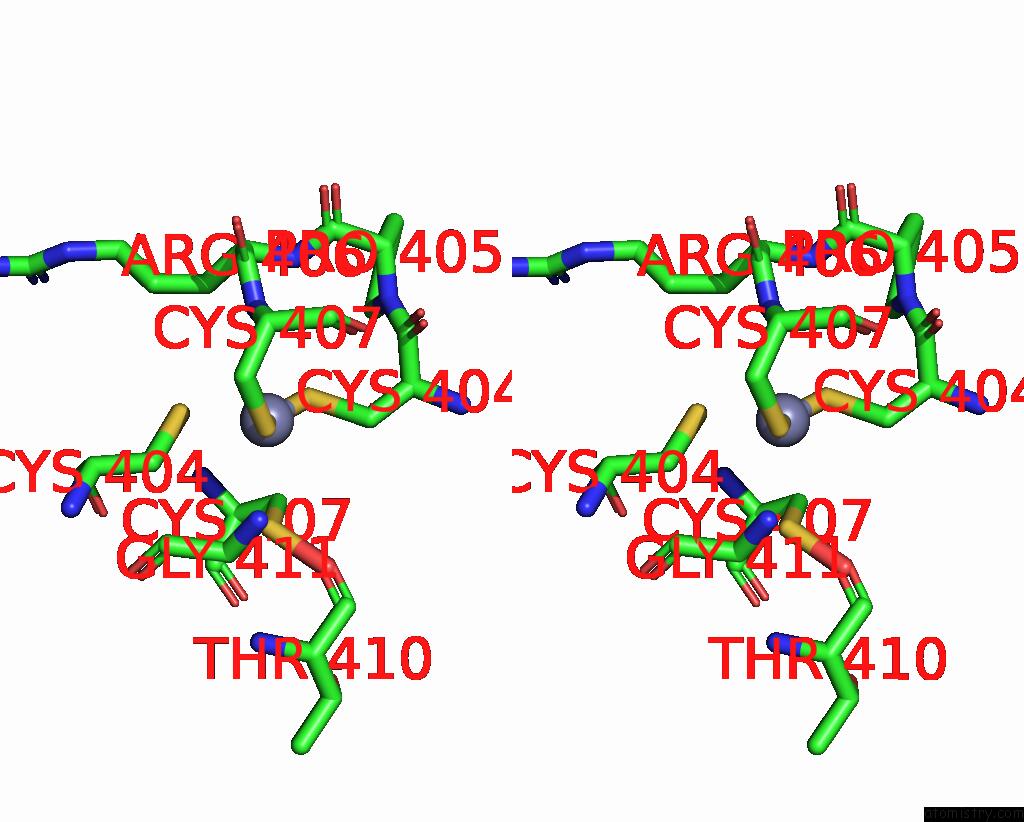
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 2 of Crystal Structure of E. Coli Rnase E Possessing M1 Rna Fragments - Catalytic Domain within 5.0Å range:
|
Reference:
D.J.Koslover,
A.J.Callaghan,
M.J.Marcaida,
E.F.Garman,
M.Martick,
W.G.Scott,
B.F.Luisi.
The Crystal Structure of the Escherichia Coli Rnase E Apoprotein and A Mechanism For Rna Degradation. Structure V. 16 1238 2008.
ISSN: ISSN 0969-2126
PubMed: 18682225
DOI: 10.1016/J.STR.2008.04.017
Page generated: Wed Aug 20 06:10:07 2025
ISSN: ISSN 0969-2126
PubMed: 18682225
DOI: 10.1016/J.STR.2008.04.017
Last articles
Zn in 3PB9Zn in 3PB8
Zn in 3PB7
Zn in 3PB6
Zn in 3PB4
Zn in 3PAN
Zn in 3PAO
Zn in 3P8B
Zn in 3PA0
Zn in 3P7W