Zinc »
PDB 5w0u-5w9s »
5w6h »
Zinc in PDB 5w6h: Crystal Structure of Bacteriophage CBA120 Tailspike Protein 4 Enzymatically Active Domain (TSP4DN, ORF213)
Protein crystallography data
The structure of Crystal Structure of Bacteriophage CBA120 Tailspike Protein 4 Enzymatically Active Domain (TSP4DN, ORF213), PDB code: 5w6h
was solved by
M.Plattner,
M.M.Shneider,
P.G.Leiman,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 49.29 / 2.29 |
| Space group | P 41 2 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 170.079, 170.079, 246.384, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 14 / 17 |
Other elements in 5w6h:
The structure of Crystal Structure of Bacteriophage CBA120 Tailspike Protein 4 Enzymatically Active Domain (TSP4DN, ORF213) also contains other interesting chemical elements:
| Potassium | (K) | 1 atom |
| Chlorine | (Cl) | 1 atom |
| Sodium | (Na) | 1 atom |
Zinc Binding Sites:
The binding sites of Zinc atom in the Crystal Structure of Bacteriophage CBA120 Tailspike Protein 4 Enzymatically Active Domain (TSP4DN, ORF213)
(pdb code 5w6h). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total only one binding site of Zinc was determined in the Crystal Structure of Bacteriophage CBA120 Tailspike Protein 4 Enzymatically Active Domain (TSP4DN, ORF213), PDB code: 5w6h:
In total only one binding site of Zinc was determined in the Crystal Structure of Bacteriophage CBA120 Tailspike Protein 4 Enzymatically Active Domain (TSP4DN, ORF213), PDB code: 5w6h:
Zinc binding site 1 out of 1 in 5w6h
Go back to
Zinc binding site 1 out
of 1 in the Crystal Structure of Bacteriophage CBA120 Tailspike Protein 4 Enzymatically Active Domain (TSP4DN, ORF213)
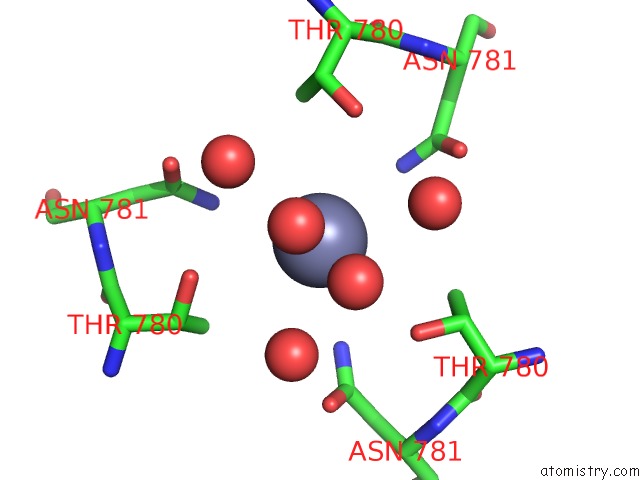
Mono view
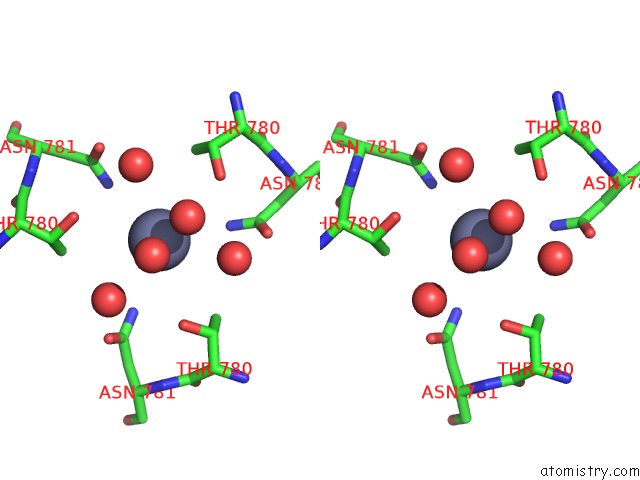
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Crystal Structure of Bacteriophage CBA120 Tailspike Protein 4 Enzymatically Active Domain (TSP4DN, ORF213) within 5.0Å range:
|
Reference:
M.Plattner,
M.M.Shneider,
N.P.Arbatsky,
A.S.Shashkov,
A.O.Chizhov,
S.Nazarov,
N.S.Prokhorov,
N.M.I.Taylor,
S.A.Buth,
M.Gambino,
Y.E.Gencay,
L.Brondsted,
E.M.Kutter,
Y.A.Knirel,
P.G.Leiman.
Structure and Function of the Branched Receptor-Binding Complex of Bacteriophage CBA120. J.Mol.Biol. V. 431 3718 2019.
ISSN: ESSN 1089-8638
PubMed: 31325442
DOI: 10.1016/J.JMB.2019.07.022
Page generated: Thu Aug 21 10:30:11 2025
ISSN: ESSN 1089-8638
PubMed: 31325442
DOI: 10.1016/J.JMB.2019.07.022
Last articles
Mn in 9LJUMn in 9LJW
Mn in 9LJS
Mn in 9LJR
Mn in 9LJT
Mn in 9LJV
Mg in 9UA2
Mg in 9R96
Mg in 9VM1
Mg in 9P01