Zinc »
PDB 3cjp-3czs »
3cww »
Zinc in PDB 3cww: Crystal Structure of Ide-Bradykinin Complex
Enzymatic activity of Crystal Structure of Ide-Bradykinin Complex
All present enzymatic activity of Crystal Structure of Ide-Bradykinin Complex:
3.4.24.56;
3.4.24.56;
Protein crystallography data
The structure of Crystal Structure of Ide-Bradykinin Complex, PDB code: 3cww
was solved by
E.Malito,
W.J.Tang,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 29.77 / 1.96 |
| Space group | P 65 |
| Cell size a, b, c (Å), α, β, γ (°) | 262.448, 262.448, 90.628, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 18.1 / 20.8 |
Zinc Binding Sites:
The binding sites of Zinc atom in the Crystal Structure of Ide-Bradykinin Complex
(pdb code 3cww). This binding sites where shown within
5.0 Angstroms radius around Zinc atom.
In total 2 binding sites of Zinc where determined in the Crystal Structure of Ide-Bradykinin Complex, PDB code: 3cww:
Jump to Zinc binding site number: 1; 2;
In total 2 binding sites of Zinc where determined in the Crystal Structure of Ide-Bradykinin Complex, PDB code: 3cww:
Jump to Zinc binding site number: 1; 2;
Zinc binding site 1 out of 2 in 3cww
Go back to
Zinc binding site 1 out
of 2 in the Crystal Structure of Ide-Bradykinin Complex
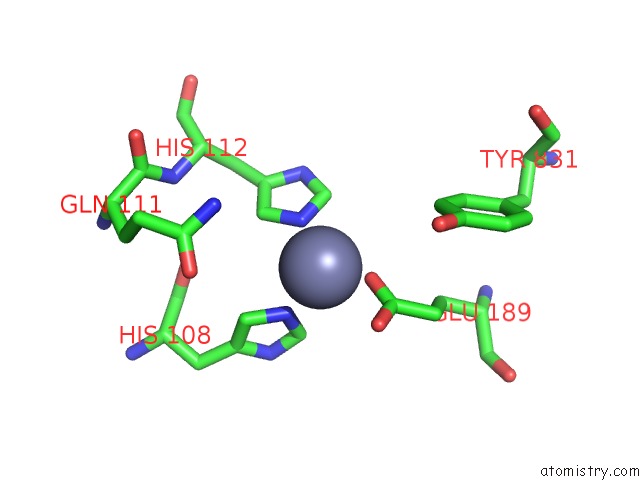
Mono view
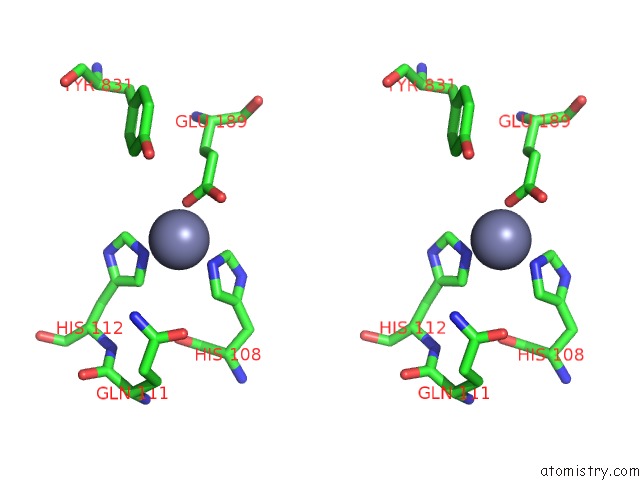
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 1 of Crystal Structure of Ide-Bradykinin Complex within 5.0Å range:
|
Zinc binding site 2 out of 2 in 3cww
Go back to
Zinc binding site 2 out
of 2 in the Crystal Structure of Ide-Bradykinin Complex
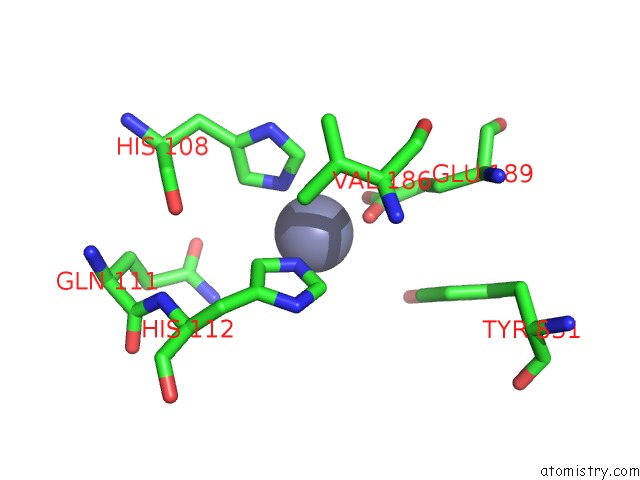
Mono view
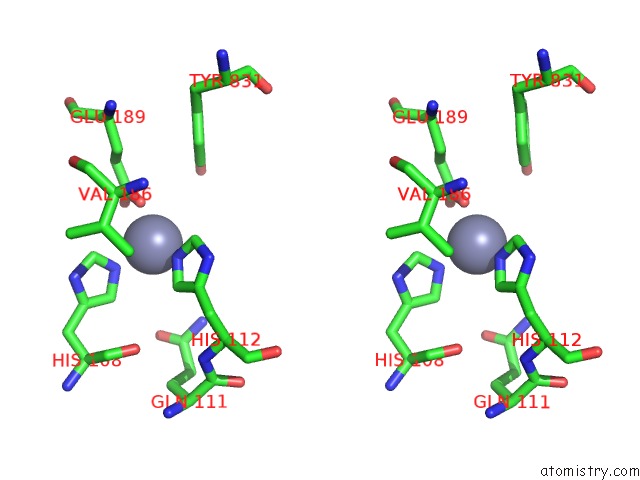
Stereo pair view

Mono view

Stereo pair view
A full contact list of Zinc with other atoms in the Zn binding
site number 2 of Crystal Structure of Ide-Bradykinin Complex within 5.0Å range:
|
Reference:
E.Malito,
L.A.Ralat,
M.Manolopoulou,
J.L.Tsay,
N.L.Wadlington,
W.J.Tang.
Molecular Bases For the Recognition of Short Peptide Substrates and Cysteine-Directed Modifications of Human Insulin-Degrading Enzyme Biochemistry V. 47 12822 2008.
ISSN: ISSN 0006-2960
PubMed: 18986166
DOI: 10.1021/BI801192H
Page generated: Thu Oct 24 11:57:51 2024
ISSN: ISSN 0006-2960
PubMed: 18986166
DOI: 10.1021/BI801192H
Last articles
K in 7G36K in 7G34
K in 7G30
K in 7G2Y
K in 7G2X
K in 7G2W
K in 7G2V
K in 7G2U
K in 7G2T
K in 7G2S